The 7-Hour Critical-Care Genome
Going from blood sample to diagnosis in the same work shift, rapid genomic sequencing proves its worth in clinical settings.
Last year, a thirteen-year-old boy in Oregon named Matthew Kunzman suffered chest pain and shortness of breath and went to see a local doctor, showing signs of myocarditis, a heart inflammation often caused by a viral infection. Kunzman’s condition was serious enough that he was airlifted to Stanford Hospital, which has a pediatric heart-transplant and mechanical-support program.
“He needed a heart and lung machine just to keep him alive,” Euan Ashley, a cardiologist at Stanford University and associate dean of the medical school, says. It was unclear if Kunzman was suffering from a treatable infection or a rare genetic disease. Doctors could perform a battery of tests, including a heart biopsy, “but it’s not a trivial procedure,” Ashley says. “You have to thread a pretty thick device through a vein into the heart, and it has little claws on the end that grab a piece of the inside of your heart.” Alternatively they could give Kunzman a genetic test. His parents selected the latter. Hours later, they definitively diagnosed the cause: Matthew indeed had a genetic condition, and three weeks later, he received a heart transplant, saving his life.
Ashley’s lab was conducting a clinical study of a new critical-care genome-sequencing method, published recently in The New England Journal of Medicine. They reported that they can go from a blood sample to diagnosis in close to seven hours, setting a Guinness World Record by halving the previous fastest time for solving a complete genome sequence. Typically, a diagnosis based on genome sequencing takes months. But, Ashley says, accelerating the process can “stop the diagnostic odyssey” and let doctors focus on treatment.
“We actually had to broaden the pipes at Stanford. We literally had to get bigger cables.”
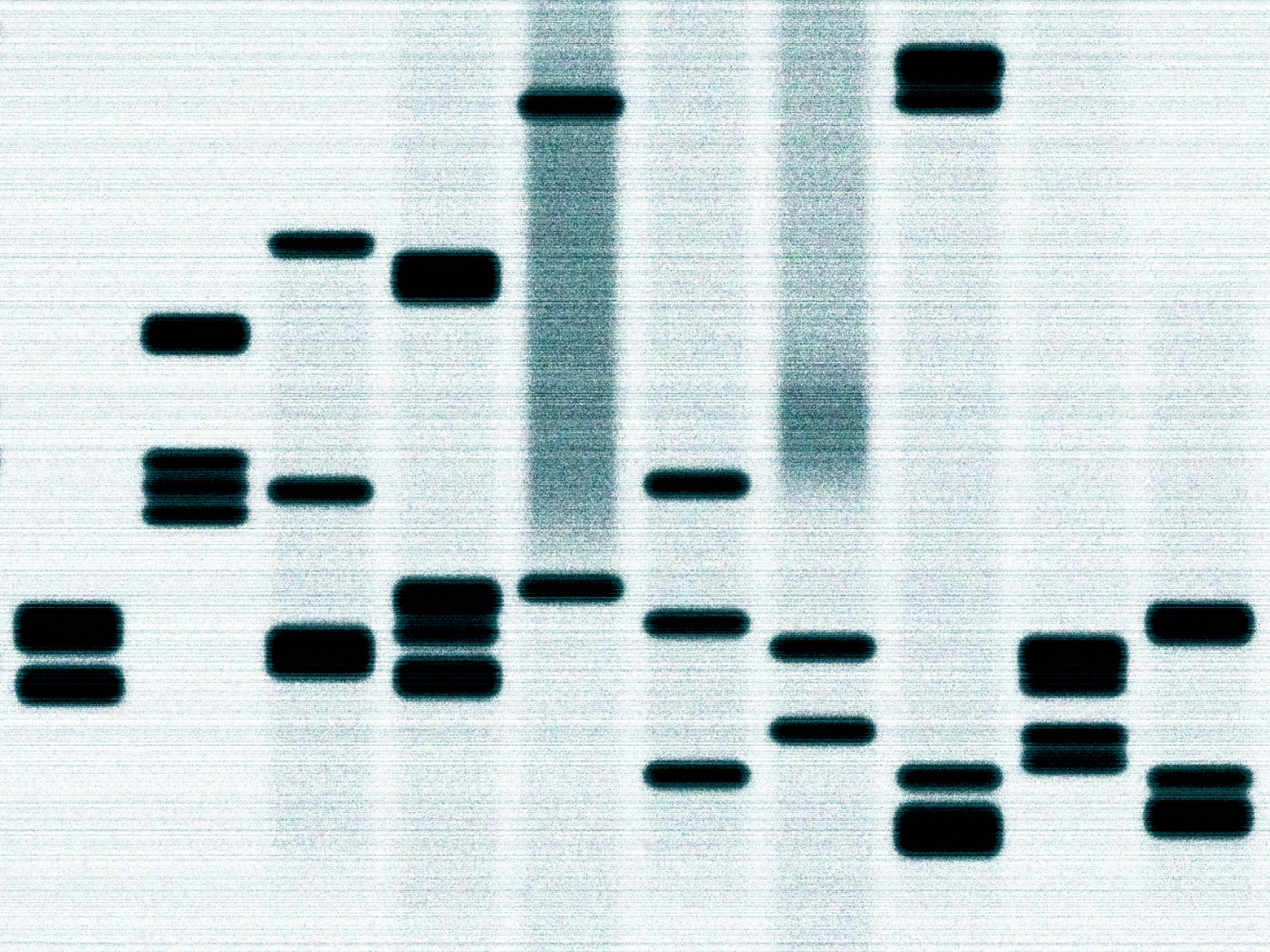
Their pipeline consists of several steps, roughly broken into three stages. In the wet-lab stage, they collect and prepare DNA samples from blood draws, then sequence them, producing an electrical amplitude signal called a “squigglegram.” In the computational stage, algorithms turn squiggles from DNA fragments into a list of bases, map them to a reference genome, find places where it varies from the reference genome, and annotate those variants. In the diagnosis stage, another algorithm filters those variants and flags any that could be relevant to the patient’s condition. Then the doctors look at the data, apply their expertise, and make a final diagnosis.
Hustle and workflow
On the face of it, there was nothing new to see here. “The different pieces of technology were available,” Ashley says. But they optimized the workflows and blew doors off with speed. First, they sequenced samples on Oxford Nanopore DNA sequencers using 48 separate “flow cells” at once. Instead of waiting for each sequence to finish and then waiting for the slow churn of a local computer to process the data, they uploaded a new batch of data every three minutes to computers in the cloud. “You can analyze it as it comes, but it comes as a tidal wave of bits,” Ashley says. “We actually had to broaden the pipes at Stanford. We literally had to get bigger cables.” Then they found a way to distribute the job over hundreds of parallel processors running on 16 computers. They avoided further delays by identifying variants in one data batch, while simultaneously identifying bases from the next batch, and sequencing the following one.
On the diagnosis side, they had to narrow roughly 4.5 million ways in which a person’s genome differs from the reference genome to just a handful of relevant genetic variant signatures. A team at the Stanford Clinical Genomics Program had constructed a complex set of filters that select variants associated with diseases, and in particular diseases whose symptoms match those of a patient like 13-year-old Matthew Kunzman.
“More than any technical thing, it was the entire team coming together,” says Sneha Goenka, an electrical engineering PhD student at Stanford and a paper co-author. It was “everyone talking to everyone, trying to understand how one stage can help the others.” Goenka found efficiencies by reducing the number of files needing to be uploaded and cleverly compressing and scheduling the rest. “It helps with the entire democratization of the pipeline,” she says. The team also made their software open source.
The researchers tested a dozen people, ranging in age from 3 months to 57 years old. All of them were suspected of having genetic diseases, including epilepsy, heart conditions, or multisystem disease for which “there isn’t really a smoking gun,” Ashley says. In the end, the team found diagnoses for five of them, the fastest in seven hours and eighteen minutes—breakneck speed for identifying a rare disease based on genomic sequencing.
“I never imagined that I would work on something so impactful.”
“It’s very exciting,” says Zornita Stark, a clinical geneticist at Victorian Clinical Genetics Services in Melbourne, who was not involved in the work. It’s “a key step towards this becoming a routine test that can be used to manage clinical care in real time.”
“I think it’s likely to be more accessible than might initially be imagined,” Ashley says, adding that they’re already looking at ways to further speed up the process, like going over a person’s genome fewer times. “We’re in an evolution towards a point where everyone will have their genome sequenced,” he says.
Oracle recently committed to investing £150 million in Oxford Nanopore, with the aim of using their own cloud services to process genomics data. “Sequence technology should be accessible in flexible settings,” says Rebecca Laborde, the senior director of Oracle Health Innovation—not just in big sequencing centers but in doctors’ offices, “when time is most vital.” She calls the Stanford work, which did not use Oracle’s software “an exciting demonstration.”
As for who will pay, eventually the cost may be negligible, but in the meantime, some countries already cover it. England’s National Health Service already considers sequencing standard care in many cases. It makes economic sense for insurance companies in the United States to also cover genomic sequencing, some experts say; reducing the cost of testing and extended hospital stays can save more than a hundred thousand dollars per patient. “We still waste a lot of time on the phone trying to persuade insurance companies to pay,” Ashley says, “but they are slowly coming around.”
You might not get a seven-hour diagnosis tomorrow. The new method “would need to go through a number of processes to be validated for diagnostic use,” Stark says. “So I think that’s still a few years away before it can be deployed at large scale.” And technology isn’t the only bottleneck. Genetic diagnosis requires “expertise to interpret the data reliably,” she says. “And we are still growing the workforce that is able to do that.”
Still, as an electrical engineer, Goenka is excited by the potential. “I never imagined that I would work on something so impactful,” she says.